Around 600,000 years ago, humanity split in two. One group stayed in Africa, evolving into us. The other struck out overland, into Asia, then Europe, becoming Homo neanderthalensis – the Neanderthals. They weren’t our ancestors, but a sister species, evolving in parallel.
Neanderthals fascinate us because of what they tell us about ourselves – who we were, and who we might have become. It’s tempting to see them in idyllic terms, living peacefully with nature and each other, like Adam and Eve in the Garden.
If so, maybe humanity’s ills – especially our territoriality, violence, wars – aren’t innate, but modern inventions.
Biology and palaeontology paint a darker picture. Far from peaceful, Neanderthals were likely skilled fighters and dangerous warriors, rivalled only by modern humans.
Top predators
Predatory land mammals are territorial, especially pack-hunters. Like lions, wolves and Homo sapiens, Neanderthals were cooperative big-game hunters. These predators, sitting atop the food chain, have few predators of their own, so overpopulation drives conflict over hunting grounds. Neanderthals faced the same problem; if other species didn’t control their numbers, conflict would have.
This territoriality has deep roots in humans. Territorial conflicts are also intense in our closest relatives, chimpanzees. Male chimps routinely gang up to attack and kill males from rival bands, a behaviour strikingly like human warfare.
This implies that cooperative aggression evolved in the common ancestor of chimps and ourselves, 7 million years ago. If so, Neanderthals will have inherited these same tendencies towards cooperative aggression.
All too human
Warfare is an intrinsic part of being human. War isn’t a modern invention, but an ancient, fundamental part of our humanity. Historically, all peoples warred. Our oldest writings are filled with war stories. Archaeology reveals ancient fortresses and battles, and sites of prehistoric massacres going back millennia.
To war is human – and Neanderthals were very like us. We’re remarkably similar in our skull and skeletal anatomy, and share 99.7 percent of our DNA.
Behaviourally, Neanderthals were astonishingly like us. They made fire, buried their dead, fashioned jewellery from seashells and animal teeth, made artwork and stone shrines. If Neanderthals shared so many of our creative instincts, they probably shared many of our destructive instincts, too.
Violent lives
The archaeological record confirms Neanderthal lives were anything but peaceful.
Neanderthalensis were skilled big game hunters, using spears to take down deer, ibex, elk, bison, even rhinos and mammoths. It defies belief to think they would have hesitated to use these weapons if their families and lands were threatened. Archaeology suggests such conflicts were commonplace.
Prehistoric warfare leaves telltale signs. A club to the head is an efficient way to kill – clubs are fast, powerful, precise weapons – so prehistoric Homo sapiens frequently show trauma to the skull. So too do Neanderthals.
Another sign of warfare is the parry fracture, a break to the lower arm caused by warding off blows. Neanderthals also show a lot of broken arms. At least one Neanderthal, from Shanidar Cave in Iraq, was impaled by a spear to the chest.
Trauma was especially common in young Neanderthal males, as were deaths. Some injuries could have been sustained in hunting, but the patterns match those predicted for a people engaged in intertribal warfare- small-scale but intense, prolonged conflict, wars dominated by guerrilla-style raids and ambushes, with rarer battles.
The Neanderthal resistance
War leaves a subtler mark in the form of territorial boundaries. The best evidence that Neanderthals not only fought but excelled at war, is that they met us and weren’t immediately overrun. Instead, for around 100,000 years, Neanderthals resisted modern human expansion.
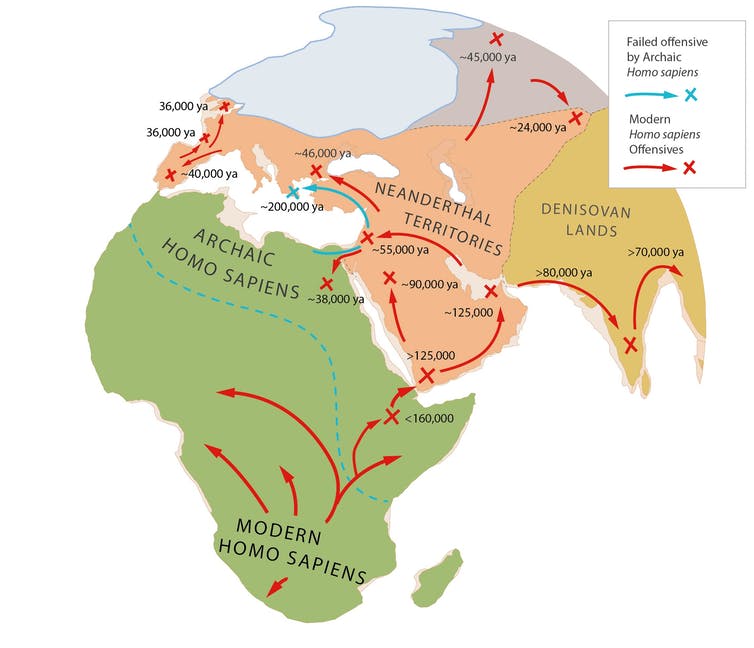
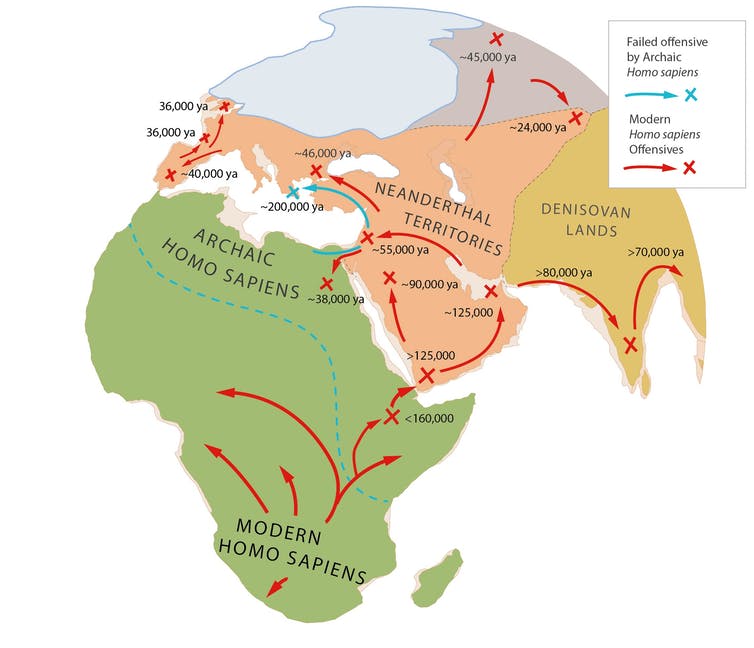
Why else would we take so long to leave Africa? Not because the environment was hostile but because Neanderthals were already thriving in Europe and Asia.
It’s exceedingly unlikely that modern humans met the Neanderthals and decided to just live and let live. If nothing else, population growth inevitably forces humans to acquire more land, to ensure sufficient territory to hunt and forage food for their children.
But an aggressive military strategy is also good evolutionary strategy.
Instead, for thousands of years, we must have tested their fighters, and for thousands of years, we kept losing. In weapons, tactics, strategy, we were fairly evenly matched.
Neanderthals probably had tactical and strategic advantages. They’d occupied the Middle East for millennia, doubtless gaining intimate knowledge of the terrain, the seasons, how to live off the native plants and animals.
In battle, their massive, muscular builds must have made them devastating fighters in close-quarters combat. Their huge eyes likely gave Neanderthals superior low-light vision, letting them manoeuvre in the dark for ambushes and dawn raids.
Sapiens victorious
Finally, the stalemate broke, and the tide shifted. We don’t know why. It’s possible the invention of superior ranged weapons – bows, spear-throwers, throwing clubs – let lightly-built Homo sapiens harass the stocky Neanderthals from a distance using hit-and-run tactics.
Or perhaps better hunting and gathering techniques let sapiens feed bigger tribes, creating numerical superiority in battle.
Even after primitive Homo sapiens broke out of Africa 200,000 years ago, it took over 150,000 years to conquer Neanderthal lands. In Israel and Greece, archaic Homo sapiens took ground only to fall back against Neanderthal counteroffensives, before a final offensive by modern Homo sapiens, starting 125,000 years ago, eliminated them.
This wasn’t a blitzkrieg, as one would expect if Neanderthals were either pacifists or inferior warriors, but a long war of attrition. Ultimately, we won. But this wasn’t because they were less inclined to fight. In the end, we likely just became better at war than they were.
Nicholas R. Longrich, Senior Lecturer in Evolutionary Biology and Paleontology, University of Bath.
This article is republished from The Conversation under a Creative Commons license. Read the original article.